Data Entry Assistant (DEA) 2.0 Procedures
Introduction
This section contains information on the practices for preparing occurrence records for entry into the xBio:D database using the Data Entry Assistant (DEA) 2.0. The DEA2 web application requires that occurrence records be present in a properly formatted data entry template (File:Data Entry Template 28-Aug-2014.xls) file according to the Data Transcription Procedures protocol or a properly pre-formatted data entry template File:DEA data entry template-full 28-Aug-2014.xls.
The DEA Processing steps do not need to be followed in the order defined in this document, but all of the parts specified do need to be completed when starting from verbatim label data. If pre-formatted specimen data is used, DEA2 will check to see if the specified values are valid according to xBio:D Controlled Vocabularies.
Contents
DEA Preparation
Login
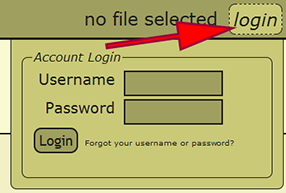
Go to the Data Entry Assistant (DEA) 2.0 web application, click on the login link in the menu on the upper-right hand part of a page, and log in. An xBio:D user account is required to prepare a file within DEA2. If an xBio:D account is needed, go to the DB Manager web application, and sign up for an account.
File Upload
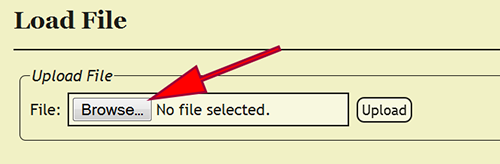
After logging in, the specimen data file, which is an Excel spreadsheet, must be uploaded into DEA2. Go to File -> Load File from the menu to go to the Load File page. Once there, click on the Browse button and select the Excel file to upload. Click the Upload button and the Excel file will be uploaded and standardized in DEA2. Standardization involves created two additional worksheets, "Main" and "Localities", and copying the specimen records from the "Raw_Data" worksheet to the "Main" worksheet, if the file has not already been standardized or pre-formatted. During standardization, DEA2 will verify the consistency of certain fields to make sure that they conform to expected values, i.e., dates are properly formatted, numbers do not contain improper characters, etc. DEA2 will also assure that each cuid is unique within the file and report those that are duplicates before proceeding. The "Main" worksheet will contain the DEA-formatted information necessary for specimen record entry into the xBio:D database. After standardization is completed, the uploaded file is set as the current loaded file and is available to begin DEA Processing.
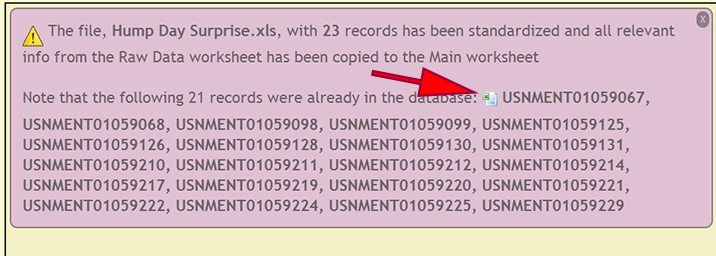
When specimen records are already within the xBio:D database, a list of cuids is displayed so the user may use the list as a reference to verify that the existing records within the database and file are authentic. There records can be downloaded within an Excel spreadsheet by clicking on the Excel logo preceding the list of records.
Note: File upload and standardization is variable and may take a few minutes to complete depending on the number of records within the Excel file.
Load Existing File
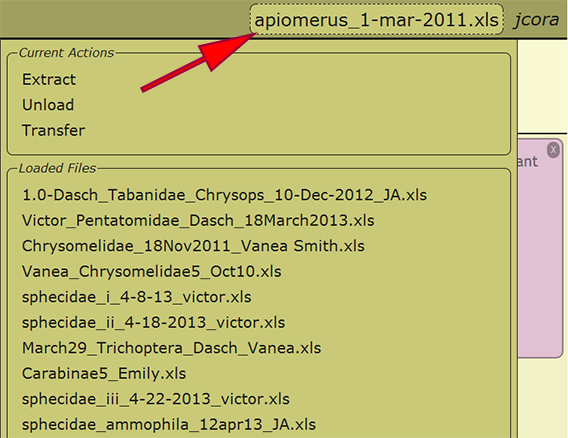
When an Excel file has been uploaded and initially loaded, the file may be reloaded from any computer in which the user has logged into DEA2. By selecting the currently loaded file or the text "no file selected" next to the username in the menu at the top right of a page, a list of the loaded files available to the user as well as some actions that may be performed on the file are presented. Any of the loaded files may be reloaded by clicking on the filename, and the recently loaded file will be displayed within the menu once loaded. Switching to a different file at any time is perfectly appropriate and will not cause any harm.
Search for File or cuid
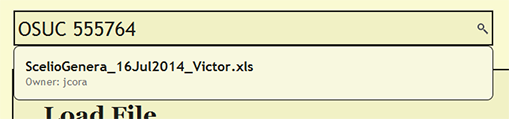
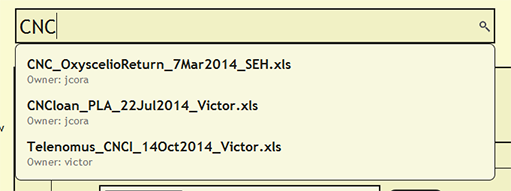
To see if a file has already been uploaded, search for the file from the search bar near the top of each page within DEA2. The search will find and list the owner of any currently loaded file that contains the search string. By searching for a cuid, the search will report the currently loaded file or files in which the cuid is present.
DEA Processing
Taxonomic Name Checking
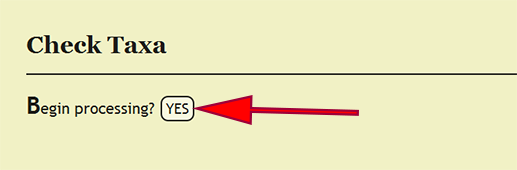
Every taxonomic name that is specified within a taxonomic column, viz., order, superfamily, family, subfamily, genus, species, and subspecies, must be present within the xBio:D database in order to enter a specimen record. Checking that these taxa are in the database is accomplished by clicking on Batch -> Check Taxa within the DEA2 menu at the top left of a page, then clicking the YES button next to Begin processing? on the Check Taxa page. DEA2 will take the taxa from the "Raw Data" worksheet, verify that they are in the database, then copy these values to their corresponding column in the "Main" worksheet. A taxonomic identification for a specimen record that is of a higher rank than family, i.e., not identified to family or more specific, will have the xBio:D database ID for that taxon placed within the family column for the specimen record. This translation will occur automatically "behind-the-scenes" during taxon checking within DEA2.
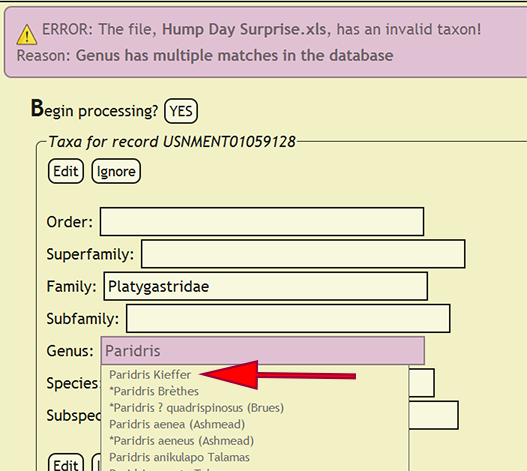
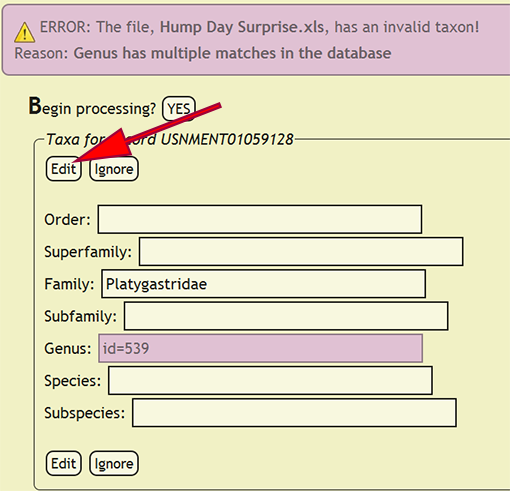
If a taxonomic name is not in the database, a form populated with the taxa for the offending specimen record will be displayed and the name that is not present will be highlighted. The form will allow the user to select the correct taxonomic name for a rank from an interactive search box, or ignore the current taxa and move on to the next group of names. The taxonomic name search box will display a list of matches from the current text string and prepend invalid taxa with an asterisk (*). If a taxonomic name has more than one taxon associated with it, i.e., homonymy, then the user can select the correct taxon with author combination to replace the homonym with its xBio:D database ID. Once the taxonomic changes have been specified, click the Edit button to reflect the modifications into DEA2 or press the Ignore button to ignore the taxa until later during specimen record entry. Batch checking of the taxa will resume once one of the two button are clicked.
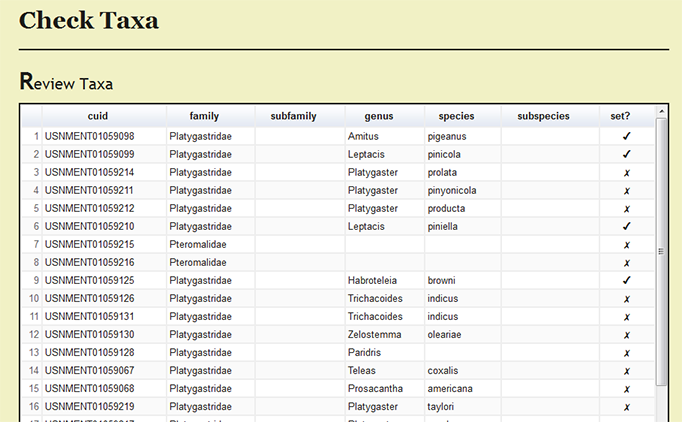
Under Page Options on the left menu, users can select Review set taxa to review which specimens records have been processed and the current taxonomic names for the records.
Users with authority over a taxonomy can enter new taxonomic names within the DB Manager web application.
Setting Determiner
Individuals within the xBio:D database are their own controlled vocabulary and are not discriminated based on whether the individual is a collector, author, or determiner. Because of this, all individuals or a collective group of individuals, known as a party, must be present within the xBio:D database. To verify that the determiners are present, go to Set -> Set Determiner from the DEA menu, which will load the Set Determiners page. The initial determiner string is the value for the record that was copied from the "Raw Data" worksheet, and this determiner name is specified within the Determiner search box. Place your cursor within the search box to interactively search for the determiner to see if his/her name is already in the xBio:D database. The use of wildcards (%) may help facilitate this search, e.g., Mues. -> Mues% -> Muesebeck, C. F. W.. Clicking on a name will copy the individual's name into Determiner box. Individual names will always be formatted with the last name (family name) first, then the person's initials. The given name parts are optional but will unambiguously identify a person when a separate individual shares the same last name and initials.
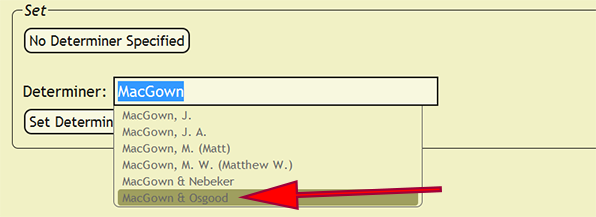
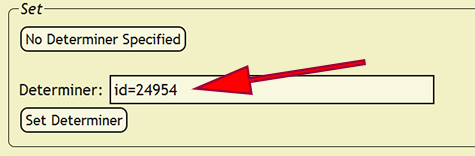
When an identification is made by more than one person, a party will need to be set as the determiner. Within the Determiner search box, search for one of the determiners, and from the list of people and parties, select the correct party. Once the party is selected, the xBio:D party ID will replace the search box text. An xBio:D person ID will also replace individual names that cannot be unambiguously specified. New parties can be formed within the DB Manager web application.
After the determiner has been confirmed and any name ambiguity removed, press Set Determiner to assign the search box text as the determiner for all of the records in which the original determiner value was present. Using the above example, Muesebeck, C. F. W. would be set as the determiner for all determiner values that matched Mues.. If a determiner is not specified, press the No Determiner Specified button to skip the current record and all matching records until the next distinct determiner value is found.
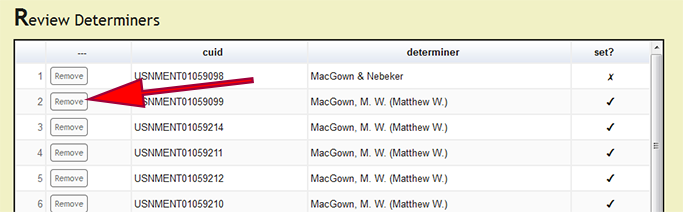
Under Page Options on the left menu, users can select Review set determiners to review which specimens records have been processed and the current determiner for the records. If a determiner was incorrectly set, press the Remove button next to the record which contains the improper determiner string to remove the action and allow the user to process the record again. DEA2 will remove the set determiner action for all of the records in which the new_comments column match with the selected record. The previously set, not original, determiner value will remain in the "Main" worksheet to allow the user to easily reset correct determiners that had their action inadvertently removed. Although this seems a bit counter-intuitive, through user feedback on real-world examples, the current behavior to remove actions for determiners is the most effective.
Changes from DEA to DEA2
- All actions performed within DEA2 are recorded, which requires an xBio:D user account. This allows a user to process a file seamlessly between multiple computers.
- DEA2 is at least 4 times faster than the original DEA, and also contains more consistency checks and better error handling.
- Verbatim label fields do not need to be merged with the comments field prior to entry into DEA2 unlike the original DEA.
- DEA2 can handle many more specimen records in a single file. Whereas DEA maxed out at ~500 specimens, DEA2 has easily processed files with as many as 8000 specimens.
- DEA2 performs some consistency checks upon upload to verify that cuids are unique within the file, dates are properly formatted, required fields are present, etc. After the consistency check, DEA2 will report any specimens within the file that are already located within the xBio:D database.